Transcription data are frequently being used to study alternative splicing in one species. However, splicing isoforms may be conserved between species of a certain evolutionary distance. Therefore, cross-species comparison of splicing isoforms may provide insight into the conservation of alternative splicing. The assessment of functional alternative splicing requires the identification of the gene product that retains the core biological function. The conservation of exonic structure between orthologous splicing isoforms of two species would suggest that they exist in both species and that their biological function may be conserved.
CExonic was developed in order to determine if splicing isoforms can be aligned to the genomic DNA of its orthologous gene in another species with conservation of exonic structure. Users can enter a VEGA identifier of a human gene or transcript and the CExonic server will align this transcript to the genomic DNA of its orthologous gene in mouse and assess whether its exonic structure is conserved in mouse. First, it determines the ortholous gene in the mouse genome, using tblastx. Then, the human transcript is aligned to this mouse genomic DNA using exonerate. Subsequently, the human and predicted mouse transcripts are aligned using muscle and the exon/intron coordinates are superimposed on the alignment, similar to exstral[1]. A schematic representation of the process can be seen in the figure below.
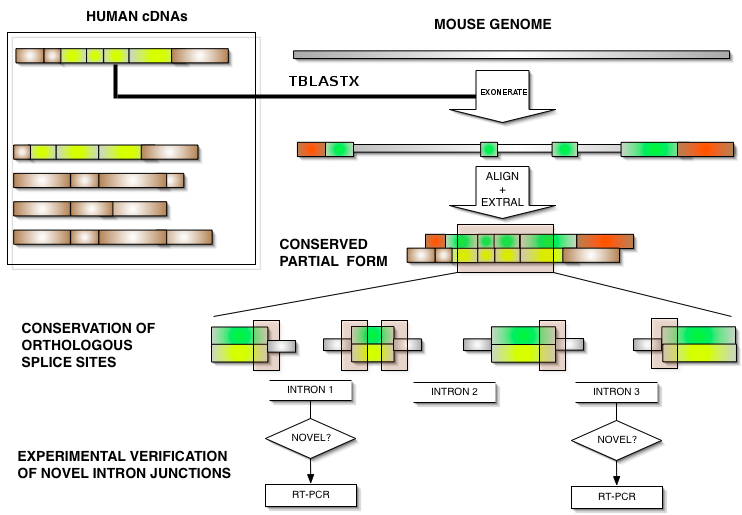
CExonic Input
CExonic can be run with human genes or transcripts for comparisons against mouse. More species will be added in the near future. Vega or EnsEMBL gene or transcript identifiers can be entered in the form, and CExonic will take their coordinates from the Vega or EnsEMBL data bases and run the comparison. Alternatively, you can specify regions of human chromosomes in the form and CExonic will then take all genes from this region for the comparison.
Pre-calculated results for human chromosome 21 and 22 can be obtained by entering VEGA human gen or transcript identifiers in the search field on the home page and pressing "enter" or the search button.
CExonic Output
CExonic generates a schematic figure of the human transcript (or gene) and of the predicted transcript in mouse. An example of these is given below, the human transcript in blue and the mouse transcript in red.
In the CExonic output, the exon/intron boundaries are represented as "><". If the aligned intron positions coincide, the exonic structure is conserved. At the end of the alignment, CExonic returns a "summary line" starting with a hash sign.
Below are a few examples of the summary line. In the first example, the transcript is conserved in mouse.
# 1 2 4 4 3
The 1 and 2 (second and third field if the # is the first) indicate sequence 1 and sequence 2, this does not change. Sequence 1 is the human sequence and sequence 2 is the mouse sequence. The name of the transcript is indicated at the beginning of the alignment. The fourth and fifth field indicate the number of exons in sequence 1 and two, respectively. The sixth field indicates the number of intron positions that could be aligned. If the number of exons in both transcripts is the same and the number of aligned intron positions is one less than the number of exons, the exonic structure is conserved.
In the example below, the exonic structure is not conserved. The sixth field indicates that no intron positions could be aligned. The fields following the sixth now indicate which intron positions could not be aligned, all four in this case.
# 1 2 5 5 0 1 2 3 4
Below the summary line, the coordinates of the non-aligned introns are given, as can be seen in the complete example of the alignment that CExonic produces below.
>OTTHUMT00000171866
1:OTTHUMT00000171866 2:OTTHUMG00000078806_mus_musculus_1
> <>
1 ATGGTTCAGCTGATTGCACCTTTAGAAGTTATGTGGAACGAGGCAGCAGATCTTAAGCC-
* *** * ******** ******** *** * * ******** * ****** *
2 -TAGTTTGTTTAATTGCACCCATAGAAGTTCTGTAGGATGAGGCAGCTGGTCTTAAATCT
> <>
1 -CCTTGCTCTGTCACGCAG--GCTGGAATGCAGTGGTGGAATCATGGCTCACTACAGCCC
** ***** * * ** ***** ** * * **** *** *** **
2 GTCTGCCTCTGCCTCCCAAGTGCTGGGAT-------TAAAGGCATGCGCCACCACACCCG
<>
1 TGACCTCCTGGGCCCAGAGATGGAGTCTCGCTATTTTGCCCAGGTTGGTCTTGAACACCT
* * * **** * **** **** * *** ** *** ** **** **
2 GCTTATTTTTATTCTTGAGACAGGGTCTTACTATGTAACCCTGGCTGGCCTGGAACTCCC
<>
1 GGCTTCAAGCAGTCCTCCTGCTTTTGGCTT---CTTGAAGTGCTTGGATTACAGTATTTC
* * ***** ** ** **** *** ***** *** ** ** ****
2 TATGTAGACCAGTCATCGTGACTTTGACTTGGCCTTGA---TCTTTATTT--AGCGTTTC
<>
1 AGTTTTATGCTCTGCAACAAGTTTGGCCATGTTGGAGGACAATCCAAAGGTCAGCAAGTT
********* ****************************** ************ ** *
2 AGTTTTATGTTCTGCAACAAGTTTGGCCATGTTGGAGGACCATCCAAAGGTCATTGAGCT
1 GGCTACTGGCGATTGGATGCTCACTCTGAAGCCAAAGTCTATTACTGTGCCCGTGGAAAT
**** **** ****** ****** * ** **** ************** *******
2 GGCTGCTGGAGATTGGGCGCTCACCCGGACGCCAGGGTCTATTACTGTGCGGATGGAAAT
<>
1 CCCCAGCTCCCCTCTGGATGATACACCCCCTGAAGACTCCATTCCTTTGGTCTTTCCAGA
********* **** * ********** ******* **************** *****
2 CCCCAGCTCACCTCGGCATGATACACCATCTGAAGATGCCATTCCTTTGGTCTTCCCAGA
<>
<>
1 ATTAGACCAGCAGCTACAGCCCCTGCCGCCTTGTCATGACTCCGAGGAATCCATGGAGGT
***** ******** ***** ******** ***** ******* ** ****** ****
2 GTTAGAGCAGCAGCTCCAGCCTCTGCCGCCGTGTCACGACTCCGTAGAGTCCATGCAGGT
<>
1 GTTCAAACAGCACTGCCAAATAGCAGAAGAATACCATGAGGTCAAAAAGGAAATCACCCT
*** ************************** ******** ******** ** *** ** *
2 GTTTAAACAGCACTGCCAAATAGCAGAAGAGTACCATGAAGTCAAAAAAGAGATCGCCTT
<>
1 GCTTGAGCAAAGGAAGAAGGAGCTCATTGCCAAGTTAGATCAGGCAGAAAAGGAGAAGGT
******* ************************** * ** ****************** *
2 GCTTGAGGAAAGGAAGAAGGAGCTCATTGCCAAGCTGGACCAGGCAGAAAAGGAGAAGCT
<>
1 GGATGCTGCTGAGCTGGTTCGGGAATTCGAGGCTCTGACGGAGGAGAATCGGACGTTGAG
*** ****** ********* ****** ******************** ****** ***
2 GGACGCTGCTCAGCTGGTTCAGGAATTTGAGGCTCTGACGGAGGAGAACCGGACGCTGAA
1 GTTGGCCCAGTCTCAATGTGTGGAACAACTGGAGAAACTTCGAATACAGTATCAGAAGAG
* ************* ***** ** ******** ** ** **** **************
2 GATGGCCCAGTCTCAGTGTGTAGAGCAACTGGAAAACCTCAGAATCCAGTATCAGAAGAG
<
1 GCAGGGCTCGTCCTAA
********* ******
2 GCAGGGCTCCTCCTAA
<
# 1 2 7 6 3 1 2 3
Flanking exons for the non-aligned introns:
# intron:1:21:29371825,29468904:+
# intron:2:21:29380109,29470081:+
# intron:3:21:29385768,29380053:+
This process is implemented as a web service, using a Taverna workflow. This workflow can be executed from this server by entering a VEGA human gene or trancript identifier in the form and pressing the "Run CExonic" button.

(http://www.biomoby.org) is a project to develop a web services architecture for bioinformatics
BioMOBY is an international research project involving biological data hosts, biological data service providers, and coders whose aim is to explore various methodologies for biological data representation, distribution, and discovery.
OBJETIVES
- Study how to address interoperability problems that are actually being faced by bioinformatics users of web-accesible resources today, and what are the factors that promote the adoption of new approaches.
- How to balance between increasing potential for interoperability and the likelihood of widespread adoption? I.e. focus upon minimizing the barriers to entry into the system, or insist upon a set of constraints that will guarantee usefulness of components of the system.
CExonic MOBY Service
The MOBY Web Service interchanges data (MOBY objects) that are XML files and the transactions are made by Simple Object Access Protocol (SOAP). This MOBY service can be executed using several tools as for example Taverna software whose workflow file can download clicking here.
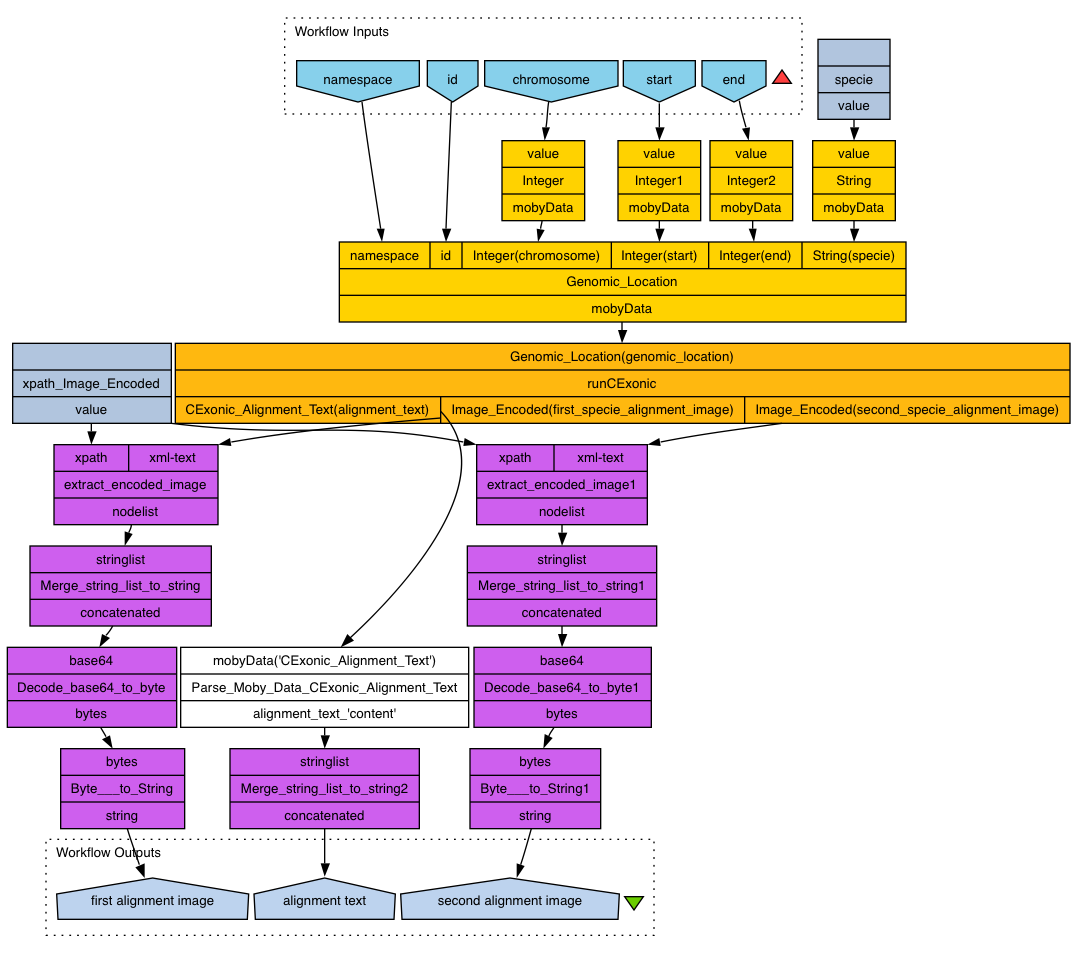

Taverna workbench (http://taverna.sourceforge.net)
Taverna allows the construction of complex in silico experiments in the form of workflows, with a particular focus on the bioinformatics domain.

Interactive Web Workflow Enactor & Manager (http://trac.bioinfo.cnio.es/trac/iwwem)
For the construction of CExonic server we have used IWWEM which is currently able to manage and enact (aka run, execute or launch) standard Taverna workflows.